Blood Transfusion Project: An AI-Oriented Model that Organizes Research into the Contents of the &psi.mRNA vial
A visual model in UML illuminating the required experiments to confirm vial contents
Sasha Latypova was writing about Graphene in the Pfizer/Moderna vials when she made this comment, appropriate here and an message to scientists everywhere on a priority experiment:
We won’t know for sure until a large quantity of them is formally seized, in multiple locations, and high quality lab analyses are made, by experts in this field of material science.
This seems obvious, but the world is running out of time to seize those vials and establish a baseline contents. Clearly, based on research from many sources, the contents will be lot dependent, and even location administered dependent. But we need this information to go to the next step, which is to model the results of introducing the vial contents into a human arm.
To guide research, as well as to establish the framework for a model that can learn from the experiments to produce a reasonable simulation of the contents introduced into a human arm, I have developed a DRAFT UML model of the vial contents. This model is based on all the available research I can get my “hands” on.
I am presenting this model in the hope that readers will contribute to it, either through validation by pointing to research papers or by adding other necessary classes, attributes, and methods.
I feel that describing the model would be too long a post, so I will present the model here and then follow with a discussion of the Classes in the model.
If you can’t read it, let me know. I’ll find a better way to present it.
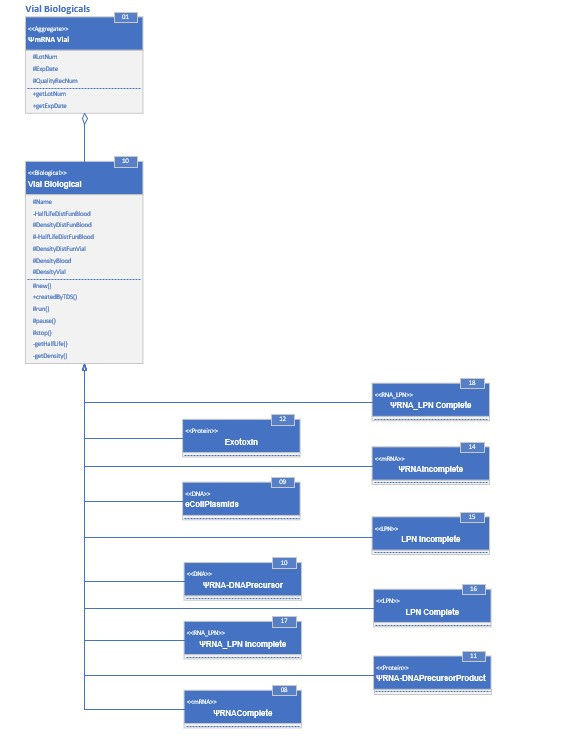
The diagram itself is “in” UML, Universal Modeling Language. You can search “UML” in YouTube to find a tutorial that suits you, or you can go to any number of web pages. Here is one: UML Tutorial. The above diagram says:
A vial class &Psi.mRNA Vial is comprised of VialBiological class. There are other classes “in” VialBiological that will be discussed later, but this is the biggie.
I have to interject here that a class is a weird thing if you are not into programming. A class is a model of a program. There is a program that runs and that asks classes to create a program- a subroutine- given that class model. That program is called an “object.” It is what runs when the model itself is running a simulation.
A bunch of classes are subclasses of VialBiological. By subclass is meant that they have all the characteristics of VialBiological plus maybe more: they inherit the characteristics of VialBiological. The “hollow” arrow indicates “inherits.”
VialBiological is represented by what we might call a “table,” which looks like a rectangle. The table has three rows. The top row contains the name of the class. In the right corner is a small rectangle containing a number. This number is used to identify a class without having to name it. So VialBiological can also be called “10”. The number will be important later on, when the model gets more complicated. Above the name in that first column is a word in « ». This is the “stereotype” of the class, which has a semantic meaning and says what “kind” of class it is. In the case of VialBiological, the stereotype is “biological”. This isn’t very informative as it stands, but it will help establish context as the full model is developed.
In the next two rows, ignore the symbols before each word in the list word list. I’ll discuss those when appropriate.
The 2nd row of VialBiological contains a list of “attributes.” These are the characteristics that will be built into any object of the class. In general, they don’t “do” something, they “are” something. They are the if and only if characteristics of the class that are of interest to the modeler. Obviously, when the class builds an object of itself, it will need to provide a “Name” to the object; and so on. I will discuss all the attributes of all the objects in subsequent posts.
The 3rd row of VialBiological contains a list of “methods.” These are, more or less, in a software sense, the functions or executables of the object built by the class. Since this is a simulation model, you can imagine that an object built by a class will have a “run()” command that will start the simulated behavior of the object as a component of the entire executing model.
I’ll continue describing this model in a later post. As a sneak peek, I will complete a model of the vial contents. That should direct scientific investigation to assemble across all research a coherent model and understanding of a vial contents (as a function of lot, location, etc). I will introduce two additional models, both models that are in the context of being “blood” models. One models all the contributions of the vial and a living cell to the blood; the other models all the contributions of a dead cell to the blood. All three models should, run as a complete model, establish the complete list of biologicals and non-biologicals to the blood as a consequence of introducing an identified vial shot to a human model. This, in turn, will establish whether or not transfusions from shot donors, either of the first two shots or of subsequent boosters, with or without natural infection, is safe for the recipient. All modeling will be time-dependent, with bayesian machine learning and assumed poisson statistics (the AI part).